CorticoMetrics FreeSurfer conversion tools
Note: FreeSurfer is a set of software tools for brain segmentation of MRI data. As such, non-brain imaging applications do not apply, and the tasks defined for DICOM4QI TID1500 component cannot be addressed. However, DICOM TID1500 Strutured Reporting is applicable, and functionality is included to encode brain region volumetric statistics. Therefore, the platform is included.
-
Description of the platform/product:
- name and version of the software:
fs2dicomv0.1.2, which works with FreeSurfer 6.0 - free?: yes; FreeSurfer is available here, and
fs2dicomis available for download from PyPI - commercial?: no
- open source?: yes; FreeSurfer and
fs2dicomsource code are hosted on GitHub - what DICOM library do you use?: DCMQI to write DICOM-SR
- name and version of the software:
-
Description of the relevant features of the platform:
fs2dicomincorporates the volumetric statistics created by FreeSurfer (saved as theaseg.statsfile) into a DICOM SR object. These can be viewed along with the DICOM SEG object in viewers such as 3D Slicer:
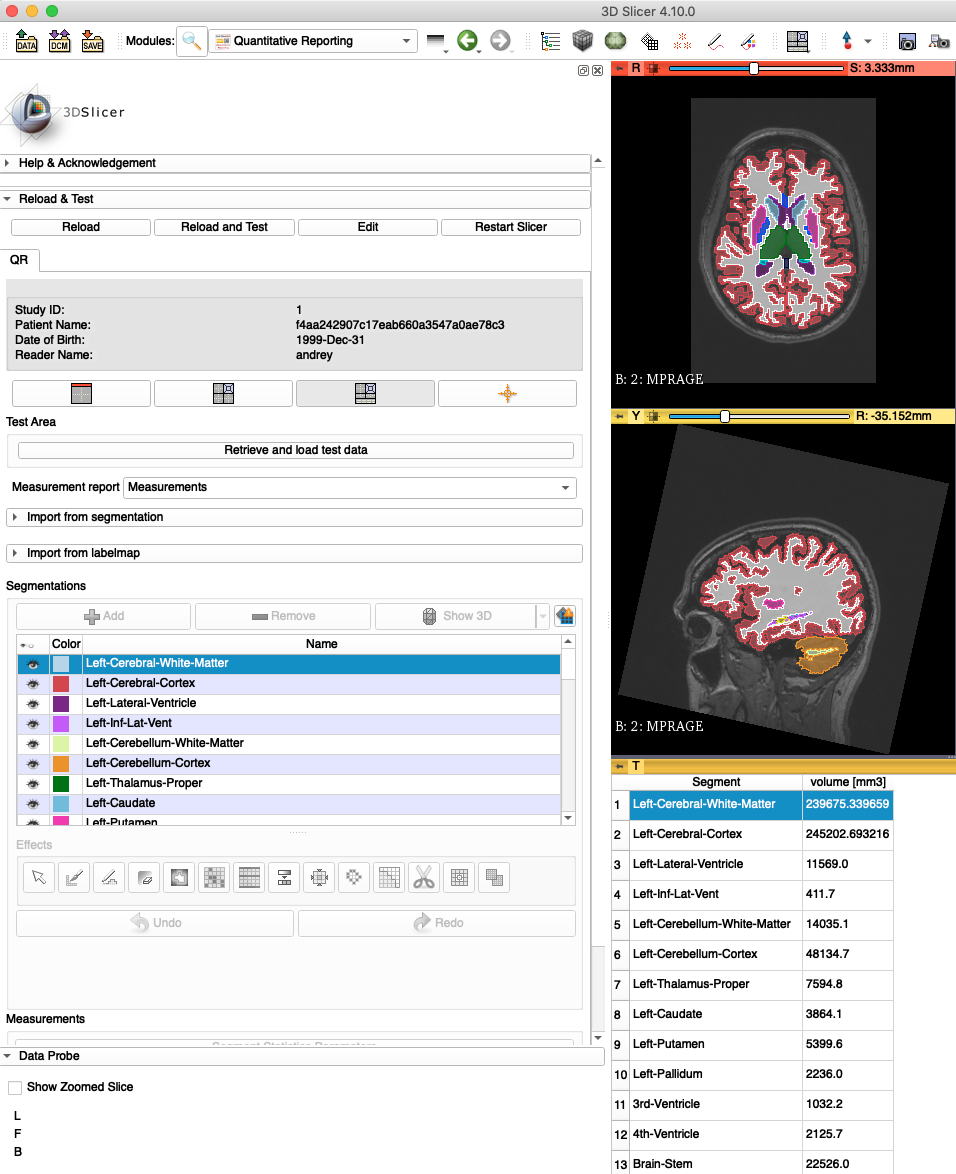
-
Read task: Not supported
-
Write task:
- Download the file
fs2dicom-rsna2018-example-v2.tar.gzby selecting the "Download -> Direct download" menu option from here. This contains the following:./dicom-anon: Directory containting input DICOMs for a FreeSurfer-compatible T1 weighted MPRAGE sequence../fs2dicom-rsna2018-example/fs-subjects/fs2dicom-rsna2018ex/: Directory containing the output of the above DICOMs processed with FreeSurfer 6.0 subcortical segmentationsfs-aseg-sr.json: File containing the volume statistics calculated by FreeSurfer, used to create the DICOM SR./aseg-sr.dcm: The DICOM SR image created byfs2dicom./aseg-seg.dcm: The DICOM SEG image created byfs2dicom, described here./readme.txt: Description of these contents, and instructions on how to runfs2dicomon the results
- Running the example (NOTE:
dockerand Python >= 3.4 are required to run this. See thereadme.txtand thefs2dicomGitHub page for more details):
pip install fs2dicom # if not installed previously tar zxvf ./fs2dicom-rsna2018-example-v2.tar.gz cd fs2dicom-rsna2018-example fs2dicom create-sr \ --aseg_dicom_seg_file ./aseg-seg.dcm \ ./dicoms-anon/MR.1.3.12.2.1107.5.2.32.35162.1999123112191238865917962 \ ./fs-subjects/fs2dicom-rsna2018ex/stats/aseg.stats \ ./aseg-sr.dcm
- Download the file